// SYSTEM_ONLINE
Aegis for
Biosecurity Screening
Detecting AI-generated malicious proteins and synthetic toxins beyond simple sequence homology.
Team Cholix — Rick Lee, Zhiyee Liang
Cholix protein interacting with its substrate NAD
// IDENTIFYING_THE_GAP
Structural security at sequence speed.
- Current sequence-level screening is easily evaded
- Expensive structure prediction tools can't scale to high-throughput screening
- Aegis infers structural context from sequence — bridging that gap
- Detects toxin-specific interface fingerprints without folding a structure
// SEQUENCE_ECOSYSTEM
A world within your sequence.
A reference fingerprint is extracted from a known toxin-substrate interface. Aegis then searches any query sequence for a match.
Residue Embeddings
Each residue gets a high-dimensional vector encoding its biochemical context.
e.g. a catalytic serine reads differently from any other serine.
Contact Maps
Pairwise distance probabilities between all residue pairs capture spatial proximity without ever predicting a 3D structure.
Interface Fingerprint
Residues with the right embeddings within spatial proximity form a unique fingerprint for a specific toxin-substrate interface.
Aegis scores any query sequence against the reference fingerprint, flagging matches regardless of sequence identity to the original toxin.
// CASE_STUDY
Case Study: Cholix Toxin
- Benchmarking Aegis against Commec (IBBIS), a sequence-based biosecurity screening tool
- Cholix toxin sequences redesigned using ProteinMPNN across a spectrum of mutational loads
- Expected benign — mutations at conserved catalytic residues
- Expected toxic — mutations outside the binding site
- 30 sequences per class per temperature (120 total) put through DNA obfuscation to diverge from wild-type
- Screened with COMMEC v1.0.0 biorisk step only — sequences matching known virulence factors flagged as Warning
| Group | n | Biorisk |
|---|---|---|
| Benign | 30 | Warning |
| Toxic | 30 | Warning |
| Group | n | Biorisk |
|---|---|---|
| Benign | 30 | Pass |
| Toxic | 15 | Warning |
| Toxic | 15 | Pass ← missed |
// CASE_STUDY
Case Study: Cholix Toxin
120 synthetic homologues of Cholix toxin were screened with Commec and Aegis. 15 toxic sequences evaded Commec entirely — Aegis flagged every one.
Blue sequences passed Commec — many still score high on Aegis
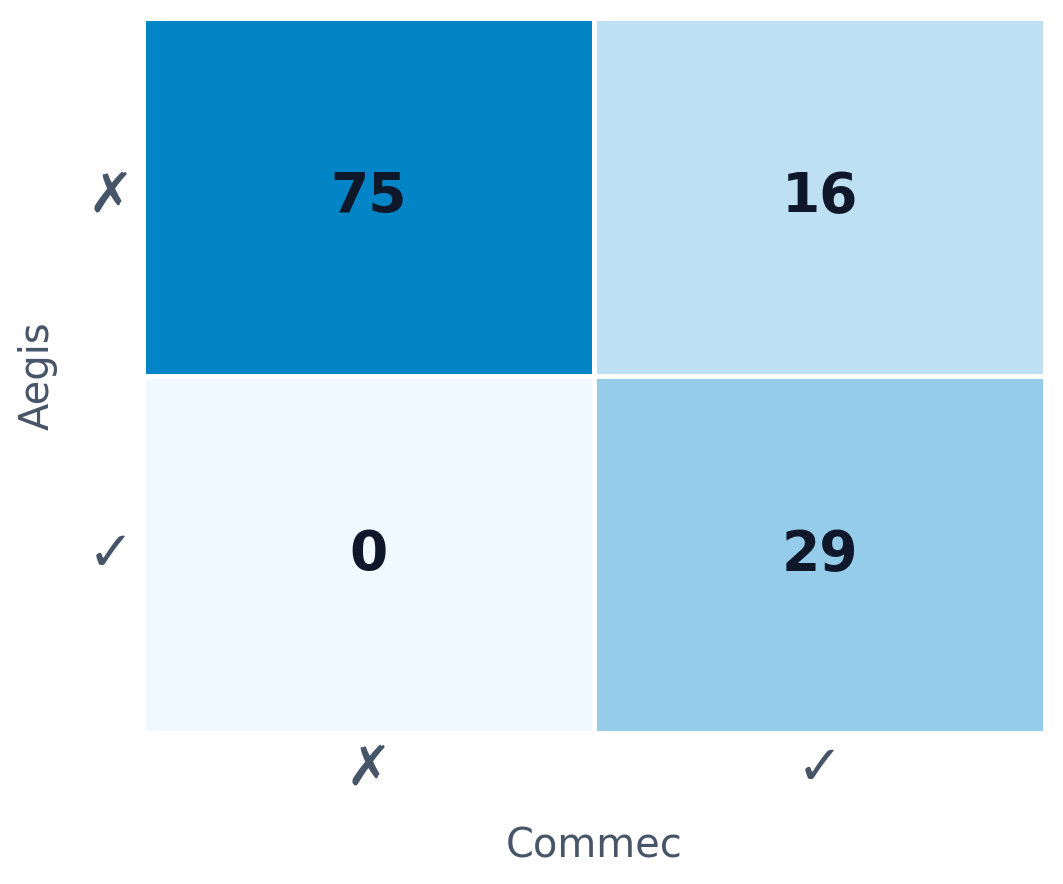
toxic sequences missed by Commec
caught by Aegis
toxic sequences missed by Aegis
// STRUCTURAL_VERIFICATION
Structural Homology: TM-score of targeted Cholix domains
3 domains: catalytic · translocation · receptor-binding
| Sequence | Max TM-score₂ | Best Target | Detection |
|---|---|---|---|
| Toxic sequence 17 | 0.859 | catalytic domain | FLAGGED |
| Toxic sequence 16 | 0.780 | catalytic domain | FLAGGED |
| Toxic sequence 4 | 0.762 | catalytic domain | FLAGGED |
| Toxic sequence 30 | 0.761 | catalytic domain | FLAGGED |
| Toxic sequence 19 | 0.749 | catalytic domain | FLAGGED |
| Toxic sequence 20 | 0.746 | catalytic domain | FLAGGED |
| Toxic sequence 29 | 0.734 | catalytic domain | FLAGGED |
| Toxic sequence 24 | 0.726 | catalytic domain | FLAGGED |
| Toxic sequence 21 | 0.724 | catalytic domain | FLAGGED |
| Toxic sequence 6 | 0.720 | catalytic domain | FLAGGED |
| Toxic sequence 9 | 0.720 | catalytic domain | FLAGGED |
| Toxic sequence 26 | 0.707 | catalytic domain | FLAGGED |
| Toxic sequence 13 | 0.699 | catalytic domain | BORDERLINE |
| Toxic sequence 25 | 0.669 | catalytic domain | BORDERLINE |
| Toxic sequence 5 | 0.652 | catalytic domain | BORDERLINE |
TM-score₂: normalised by target domain length
The 15 sequences flagged by Aegis share a similar fold with the catalytic domain of Cholix toxin, and are therefore likely to be toxic.